ERC Develops Computational-Experimental Strategy for Generating Novel Catalytic Capabilities
Researchers at the Center for Biorenewable Chemicals (CBiRC), an NSF-funded Engineering Research Center (ERC) headquartered at Iowa State University (ISU), have developed a heuristic strategy, known as CompEx, for redesigning biocatalysts and generating novel catalytic capabilities. Based on the combined efforts of CBiRC’s Thrust 1 team, the strategy integrates both experimental and computational modeling approaches to visualize the 3-D structure of active site pockets of enzymes.
This capability enables formulation of testable hypotheses on how to redesign an active site pocket to accept alternative substrates, thereby establishing new metabolic capabilities. Understanding the design principles of an active site pocket of enzymes has tremendous potential in redesigning metabolic processes to generate novel biochemicals that are not accessible in biological systems. CompEx thus provides CBiRC scientists and the broader biocatalytic community with formalized capabilities to alter and redesign metabolic processes and target the synthesis of specific end-products. CBiRC’s Thrust 1 team illustrated application of the CompEx strategy by several redesigns, which are innovating technologies in CBiRC’s three testbeds.
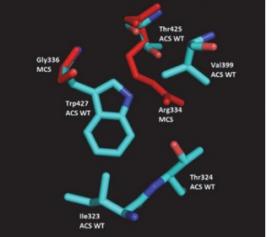
CompEx, by its use of a combined COMPutational-EXperimental approach, provides a robust strategy for taking advantage of the available 3-D structures. It predicts how one can manipulate the primary sequence of a protein to induce designed effects on catalytic specificity. The several redesigns accomplished by CbiRC's Thrust 1 team are pyrone synthase(s) (Project T1.9), acyl-ACP thioesterases (Project T1.6), and acyl-CoA synthetases (Project T1.3) (see figure).
Manipulating the substrate specificity of enzymes has traditionally been an empirical approach relying heavily on experimental data concerning the 3-D structure of enzymes. Current capabilities for experimentally determining 3-D structures are considerably slower than the rate at which primary structures are determined. However, the number of 3-D structures that are available in the PDB database (more than 105 structures) has asymptotically reached a maximum number of protein folds (about 103), indicating that biology has evolved a large number of biocatalysts on a limited number of 3-D scaffolds. Therefore, subtle chemo-physical variations on the catalytic surface of enzymes guide the evolution and development of enzyme diversity. Moreover, because these enzymatic catalytic surfaces are dynamic during the catalytic cycle, chemico-physical variations at a distance from the catalytic surface can also affect catalysis and catalytic rate constants.